Deep learning and web technologies are the focus of 3D Slicer 2017 Winter Project Week

Earlier this month, we attended the 3D Slicer 2017 Winter Project Week at the Massachusetts Institute of Technology (MIT) Computer Science and Artificial Intelligence Laboratory (CSAIL) in Cambridge, MA. The project week is a semi-annual event, where 3D Slicer users and developers gather to discuss and work collaboratively on open-science solutions for problems that occur at the intersection of the computer science, mechanical engineering, biomedical engineering, and medical fields.
During the week, we joined approximately 100 attendees—mainly from the U.S., with some traveling from Canada, China, Denmark, Germany, Italy, Netherlands, Norway, Spain, and Sweden—to work on close to 60 projects that cover deep learning and graphics processing units (GPUs), web technologies, image-guided therapy (IGT), diffusion magnetic resonance imaging (MRI), quantitative imaging informatics, shape analysis, segmentation, 3D Slicer infrastructure, 3D Slicer training, and 3D Slicer dissemination. We made progress on specific technical issues; helped the 3D Slicer community with our expertise; and discussed 3D Slicer’s design, roadmap, and key priorities.

Overall, the consensus at the project week was to continue to maintain a lean, robust, and well-documented 3D Slicer core that can be easily extended and to enable the use of the platform in the cloud for easier integration with web technologies, cloud-based infrastructures and deep learning frameworks like Tensorflow.
Additional highlights are as follows.
Web-technologies Breakout Session
This breakout session provided an interesting showcase of the different web-technology approaches followed by the 3D Slicer community. Different groups presented their work on data federation, remote integration, and cloud processing using technologies such as JavaScript, HTML5 or WeGL.
Girder Breakout Session
Girder is a web platform for medical data management and processing developed at Kitware. After learning more about existing use cases (e.g., DCMQI, Chronicle, and Arbor). Brian Helba and Jean-Christophe Fillion-Robin presented the overall Girder platform and demoed the Girder-based project HistomicsTk. HistomicsTk enables the visualization of large titled microscopy images along with the remote execution of 3D Slicer command-line interfaces for processing. This session helped the community understand the flexibility of the Girder platform and how it can be leveraged, extended, and integrated. This organization of this session was supported by National Cancer Institute (NCI) award 1U24CA194354.
Shape Analysis Breakout Session
A recently funded large National Institutes of Health (NIH) project led by Kitware will greatly enhance 3D Slicer’s functionality with novel, state-of-the-art shape analysis methodologies (see figure below). This breakout session led by Beatriz Paniagua was intended to allow the project team to synchronize with the 3D Slicer community to agree on architectural design choices to move forward with the project.
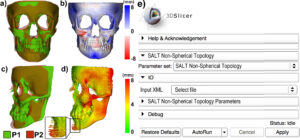
Volumetric Mesh Support
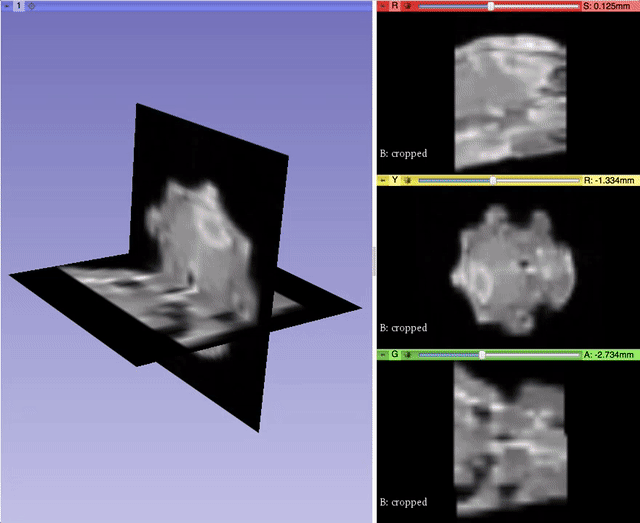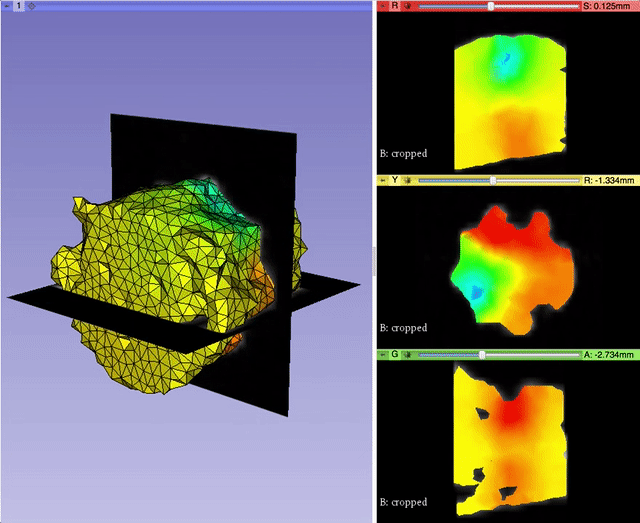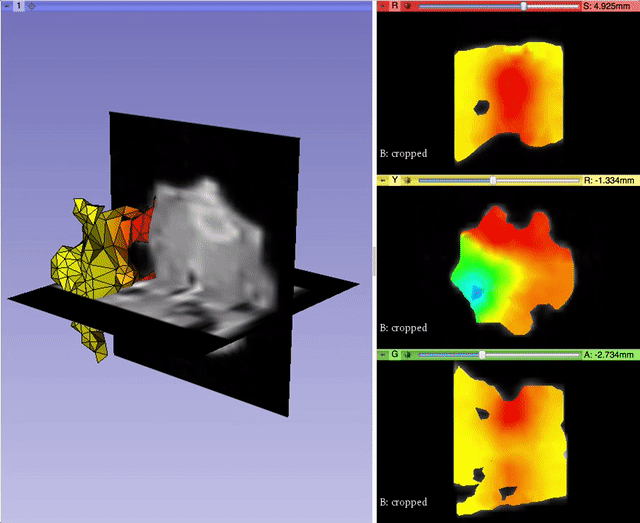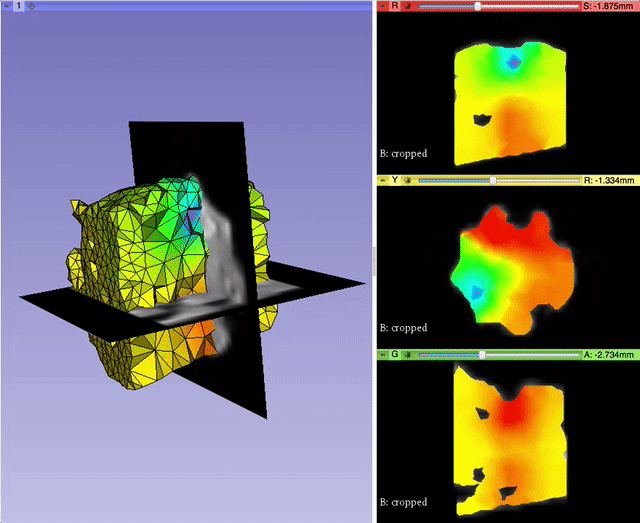
Alexis Girault from Kitware led the effort to bring back volumetric mesh visualization capabilities within 3D Slicer, a work initiated by Curtis Lisle and Steve Pieper in the previous major release of 3D Slicer. A pull request is currently under review. It will enable 3D Slicer to read/write and display finite element volumetric meshes within its three-dimensional and slice viewers. By taking advantage of the modularity of VTK data structures, Alexis was able to generalize the Model infrastructure to work with vtkPointSet, the base class of vtkPolyData (surface mesh) and vtkUnstructuredGrid (volumetric mesh). With the help of Andras Lasso and the feedbacks of numerous other Slicer developers, Alexis refactored the visualization pipeline to better manage displaying models, and improved the module interface for usability. This work was partially supported by National Cancer Institute (NCI) award R01CA184354 as part of NIRFAST-Slicer improvements. More details will be outlined in a dedicated follow-up blog post.
Qt5 Support
Jean-Christophe Fillion-Robin from Kitware led the effort to add support for Qt5. In addition of updating Slicer to support configuration using either Qt4 and Qt5, this work also includes the update of dependent project like CTK and VTK. A preliminary pull request is ready for review. This work was partially supported by National Cancer Institute (NCI) award 1U24CA194354.
Improved Transform Interaction
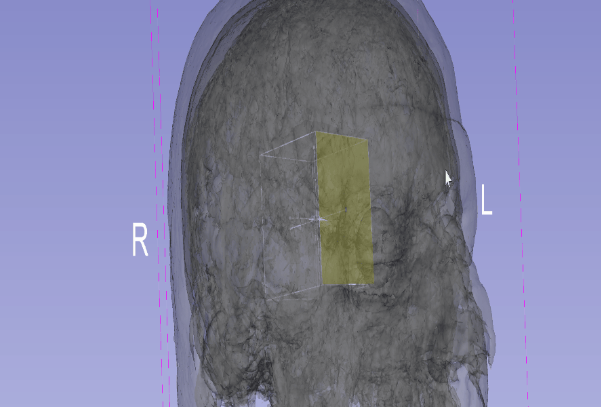
Johan Andruejol from Kitware led the effort to improve 2D and 3D view user interaction by supporting interactive translation, rotation and scaling of models and images. After creating a proof of concept, Johan had a good basis to discuss with Andras Lasso from Queens and clearly define the widget allowing user to interact. It was decided that the transformations widget should automatically be centered around the objects to transform. Support will be first added in the 3D view and then in the 2D view. This work was partially supported by the NIH award R42-HD081712 part of the Craniosynostosis effort.
Project Week Tutorial Contest
This year, Kitware was proud to award $250 to the project week tutorial contest winner, Csaba Pinter from Queen’s University, who explained how to use the segmentation module for 3D printing. More details will be outlined in a dedicated follow-up blog post.
To wrap up this post, we will again leave you with some food for thought. 3D Slicer is no longer a monolithic infrastructure; it is a modular framework that allows you to either create professional-scale applications or quickly prototype ideas. Similar to what ParaView is for scientific computing, 3D Slicer is your tool for medical imaging. Both ParaView and 3D Slicer allow you to share your creative and efficient solutions built on top of CMake, the Common Toolkit (CTK), the Insight Segmentation and Registration Toolkit (ITK), Python, or the Visualization Toolkit (VTK).
If you would like help adapting 3D Slicer to your projects, please contact Jean-Christophe Fillion-Robin via kitware@kitware.com.